Neural network for malaria detection on segmented cell images. The current state of the project is summarized here and I will update this as further work is done.
- Python 3
- PyTorch
- Matplotlib
- Pandas (for results analysis)
- Numpy (for results analysis)
The dataset used for this project is provided by the National Institutes of Health. Available here.
The dataset consists of 27,558 segmented cells from the thin blood smear slide images of 151 patients used for malaria screening. Cells are "stained" with an agent to distinguish the parasite. The dataset contains an equal number of parasitized and uninfected specimens.
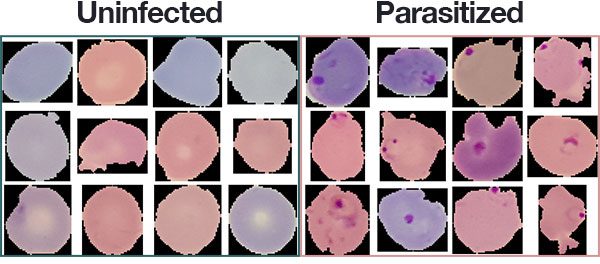
Healthy cells usually display a uniform appearance whereas parasitized cells will display distinctive marks (image courtesy of pyimagesearch):
Dataset was divided into three splits : training (60%), validation(20%) and test (20%).
Images are resized to 96x96 before training, normalized by channel and augmented with rotation, translation, sheering, scaling and flipping.
All models are defined in models.py. The PaperCNN model is detailed below:
Two other models were tested so far. The 'CustomVGG' models follows a similar architecture, but stacks additional convolutional layers before pooling and relies on 3x3 filters [Input > 2 x Conv(3x3) > Max-Pool + BN > 3 x Conv(3x3) > Max-Pool + BN > 3 x Conv(3x3) > Max-Pool + BN > Avg-Pool > FC(1x256) > Output].
Finally, the 'CustomInception' model applies a single 3x3 convolution and a stack of two 3x3 convolutions, both on the same input. The output of both branches are concatenated on the channel dimension and pooled before moving forward. The process is repeated three times before average pooling and sending to the fully-connected layer detailed above. The idea was to see if having two different receptive fields working on the same input in parallel would lead to any improvements.
ReLU was used for all activations.
The best results achieved so far with the three models are the following :
| Accuracy | PaperCNN | CustomVGG | CustomInception |
|---|---|---|---|
| Train | 98.35 % | 99.63 % | 98.76 % |
| Validation | 97.70 % | 98.08 % | 97.55 % |
And here's a summary of the best experiment (98.08 % on validation set):
| Model | Optimizer | Epochs completed | Patience | Learning Rate | Dropout (convolutions) | Dropout (fully connected) |
|---|---|---|---|---|---|---|
| CustomVGG | Adam | 536 | 200 | 0.001 | 0.05 | 0.5 |
I will provide results on the test set after implementing the changes detailed below.
Results do not quite match those achieved with similar models by Rajaraman and al. (99.09 % with their custom CNN). Reading the paper gave a few ideas as to how to improve this project:
- Cell images were randomly assigned to training, validation and test sets. This should arguably have been done with patients instead (there are multiple cell specimens per patient) to avoid any 'data leakage' and contamination of the validation and test sets.
- Models could probably benefit from pretraining on a larger dataset to strengthen the extraction of generic features (e.g. ImageNet).
- Hyper-parameters were tuned using basic grid-search on handpicked discrete values. The process should be randomized and applied on ranges instead.
- Models should be evaluated using k-fold cross-validation and other performance metrics.