Comments (11)
I'm not sure what you're referring to by "base" in this context. It sounds like you want something along the lines of this example: https://pylidc.github.io/tuts/consensus.html ?
For example,
import numpy as np
import matplotlib.pyplot as plt
import pylidc as pl
from pylidc.utils import consensus
# Query for a scan, and convert it to an array volume.
scan = pl.query(pl.Scan).filter(pl.Scan.patient_id == 'LIDC-IDRI-0001').first()
vol = scan.to_volume()
nodules = scan.cluster_annotations()
annotations = nodules[0]
consensus_mask, consensus_bbox, _ = consensus(
annotations,
clevel=0.5,
pad=[(20,20), (20,20), (0,0)]
)
k = consensus_mask.shape[-1] // 2
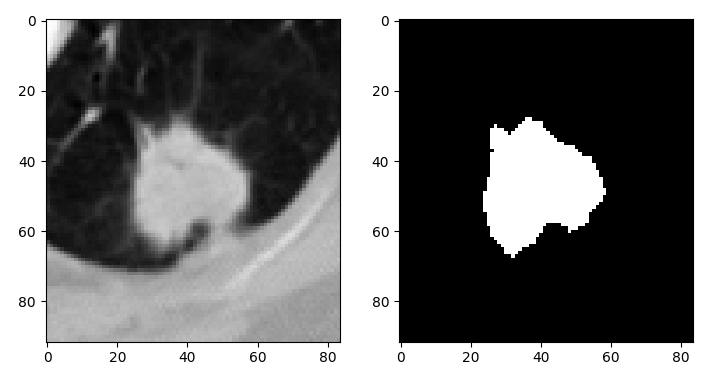
plt.subplot(121)
plt.imshow(vol[consensus_bbox][:, :, k], cmap='gray')
plt.subplot(122)
plt.imshow(consensus_mask[:, :, k], cmap='gray')
plt.show()Outputs:
from pylidc.
I wasn't very clear. I mean, to save each annotations from dataset LIDC as a new image. To save just the annotation region. Thanks!
from pylidc.
So, is what I posted above what you're looking for or...?
from pylidc.
I'll try it. I guess it will works. If don't work I post here, if work I'll post too. Thanks one more time.
from pylidc.
OK. Feel free to close this issue if you find the above examples work for you.
from pylidc.
Hi, your post was useful. Can help me in more one thing? Is possible save just nodules of GGO and consolidation, for exemple?
from pylidc.
Yes, but keep in mind that it depends on what you mean by a GGO nodule, since attributes like GGO are assigned individually by the annotators. So, do you mean nodules where at least 1 annotator assigned GGO as the texture attribute? At least 50% of annotators?
In any case, I would start by looping through each Scan, and cluster the annotations using Scan.cluster_annotations in order to group annotations belonging to the same physical nodule. So, something like:
for scan in pl.query(pl.Scan):
# annotation_groups is a list of of lists of Annotation's
annotation_groups = scan.cluster_annotations()Next, for each annotation group, implement your criteria of what qualifies as GGO. E.g.,
for nodule_annotations in annotation_group:
# Only consider nodules with 4 annotators and have >= 50% indicating GGO
if (len(nodule_annotations) == 4 and
sum([a.texture == 1 for a in nodule_annotations]) >= 2):
# Do whatever you want, e.g.,
# consensus(nodule_annotations)from pylidc.
Thanks! It works.
from pylidc.
Man, I have another issue. I got do this for 2D nodules images. How can I do the same for 3D volume of interest? Thanks!
from pylidc.
How can I do the same for 3D volume of interest?
The image and consensus masks are volumes.
from pylidc.
Ok. I did this:
for scan in pl.query(pl.Scan):
annotation_groups = scan.cluster_annotations()
vol = scan.to_volume()
for nodule_annotations in annotation_groups:
if (len(nodule_annotations) >= 2 and sum([a.texture == 1 for a in nodule_annotations]) >= 1):
consensus_mask, consensus_bbox, _ = consensus(
nodule_annotations,
clevel=0.5,
pad=[(5,5), (5,5), (0,0)]
)
image = np.asarray(vol[consensus_bbox][:, :, :]).transpose(2,0,1)
mask_image = np.float32(np.array(consensus_mask[:, :, :])).transpose(2,0,1)Thanks!
from pylidc.
Related Issues (20)
- How to access NBIA via pylidc HOT 1
- ValueError: The length of the pixel data in the dataset (524285 bytes) doesn't match the expected length (524288 bytes). HOT 5
- how to obtain the physical bbox? HOT 5
- Where to find pylidc.conf file in Google Colab? HOT 1
- Not able to do path configuration for pylidc on mac HOT 1
- RuntimeError: Could not establish path to dicom files. Have you specified the `path` option in the configuration file C:\Users\user\pylidc.conf? HOT 1
- marching_cubes_lewiner HOT 1
- Pylidc Annotation Class HOT 3
- 3D Annotation HOT 9
- OSError: Couldn't find DICOM files for Scan(id=12,patient_id=LIDC-IDRI-0001). HOT 6
- Showing patients with multiple nodules HOT 9
- module 'cmap_d' error
- RuntimeError: Could not establish path to dicom files in MacOS HOT 6
- Mismatching SeriesInstanceUID and StudyInstanceUID HOT 2
- Using populate.py
- AssertionError: Internal structure score out of bounds in Annotation(id=3761,scan_id=516)
- Annotation slice number HOT 2
- centroid to physical coordinates HOT 1
- Majorly slow dataloader using pylidc HOT 1
- Fix deprecated numpy dtype
Recommend Projects
-
 React
React
A declarative, efficient, and flexible JavaScript library for building user interfaces.
-
Vue.js
🖖 Vue.js is a progressive, incrementally-adoptable JavaScript framework for building UI on the web.
-
 Typescript
Typescript
TypeScript is a superset of JavaScript that compiles to clean JavaScript output.
-
TensorFlow
An Open Source Machine Learning Framework for Everyone
-
Django
The Web framework for perfectionists with deadlines.
-
Laravel
A PHP framework for web artisans
-
D3
Bring data to life with SVG, Canvas and HTML. 📊📈🎉
-
Recommend Topics
-
javascript
JavaScript (JS) is a lightweight interpreted programming language with first-class functions.
-
web
Some thing interesting about web. New door for the world.
-
server
A server is a program made to process requests and deliver data to clients.
-
Machine learning
Machine learning is a way of modeling and interpreting data that allows a piece of software to respond intelligently.
-
Visualization
Some thing interesting about visualization, use data art
-
Game
Some thing interesting about game, make everyone happy.
Recommend Org
-
Facebook
We are working to build community through open source technology. NB: members must have two-factor auth.
-
Microsoft
Open source projects and samples from Microsoft.
-
Google
Google ❤️ Open Source for everyone.
-
Alibaba
Alibaba Open Source for everyone
-
D3
Data-Driven Documents codes.
-
Tencent
China tencent open source team.

from pylidc.