Python module to make publication quality plots of weighted, directed graphs of medium size (10-100 nodes). Unweighted, undirected graphs will look perfectly fine, too. The node positions can be tweaked using the mouse (after an initial draw). It only depends on numpy, scipy, and matplotlib.
Existing draw routines for networks/graphs in python (networkx, igraph) use fundamentally different length units for different plot elements. This makes it hard to
- provide a consistent layout for different axis / figure dimensions, and
- judge the relative sizes of elements a priori.
This module amends these issues.
Furthermore, algorithmically finding a visually pleasing layout of
node positions is, in general, difficult. This is demonstrated by the
plethora of different algorithms in use (if graph layout was a solved
problem, there would only be one algorithm). To ameliorate this
problem, this module contains an InteractiveGraph class, which allows
node positions to be tweaked with the mouse (after an initial draw).
import numpy as np
import matplotlib.pyplot as plt; plt.ion()
import netgraph
# Construct sparse, directed, weighted graph
# with positive and negative edges:
total_nodes = 20
weights = np.random.randn(total_nodes, total_nodes)
connection_probability = 0.2
is_connected = np.random.rand(total_nodes, total_nodes) <= connection_probability
graph = np.zeros((total_nodes, total_nodes))
graph[is_connected] = weights[is_connected]
# Make a standard plot:
netgraph.draw(graph)
# Create an interactive plot.
# NOTE: you must retain a reference to the object instance!
# Otherwise the whole thing will be garbage collected after the initial draw
# and you won't be able to move the plot elements around.
plot_instance = netgraph.InteractiveGraph(graph)
# Access new node positions:
pos = plot_instance.node_positionsnetgraph.draw supports various formats for the graph argument.
In order of precedence:
-
Edge list:
Iterable of (source, target) or (source, target, weight) tuples, or equivalent (m, 2) or (m, 3) ndarray.
-
Adjacency matrix:
Full-rank (n,n) ndarray, where n corresponds to the number of nodes. The absence of a connection is indicated by a zero.
-
igraph.Graph or networkx.Graph object
import networkx
g = networkx.from_numpy_array(graph, networkx.DiGraph)
netgraph.draw(g)There are many ways to customize the layout of your graph. For a full list of available arguments, please refer to the documentation of
drawdraw_nodesdraw_edgesdraw_node_labelsdraw_edge_labels
Easiest via pip:
pip install netgraph
For the newest and brightest (and probably buggiest) version:
pip install git+https://github.com/paulbrodersen/netgraph.git
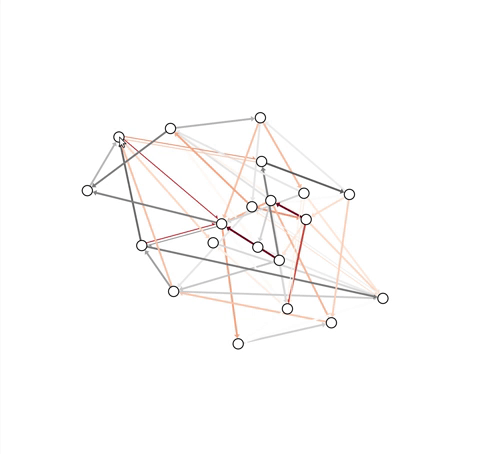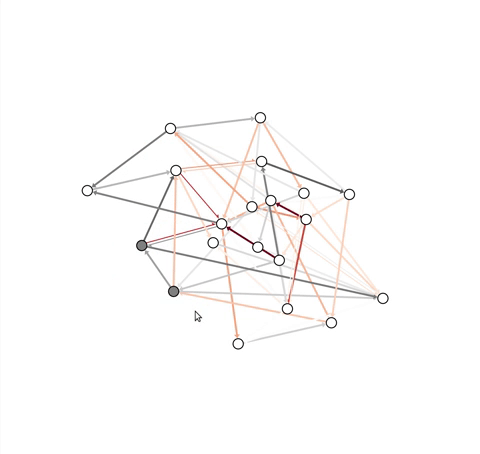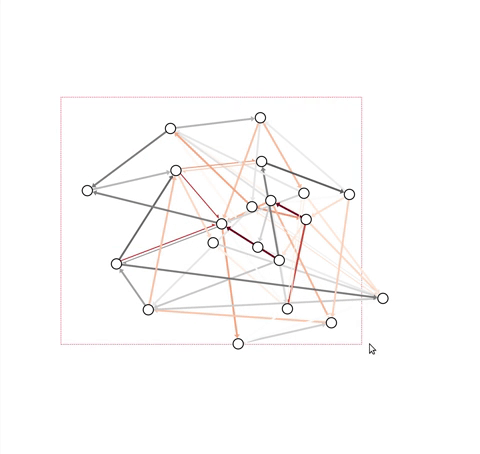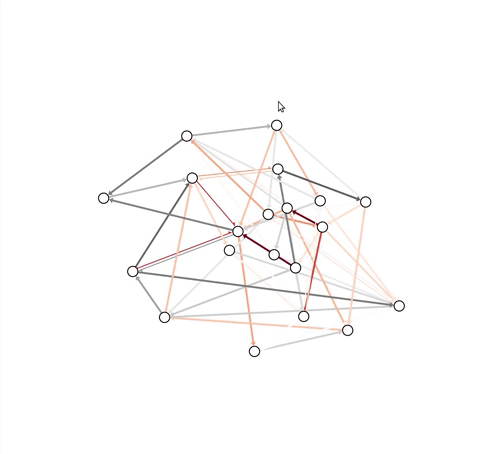
The following images show the netgraph output when using the default
settings, i.e. the output of draw in the absence of any arguments
other than graph.
Default plot for a directed, weighted network:

No arrows are drawn if the network appears undirected:

Edge weights are mapped to edge colors using a diverging colormap, by default 'RdGy'.
Negative weights are shown in red, positve weights are shown in gray.
A directed network with purely positive weights hence looks like this:

Unweighted networks are drawn with uniformly black edges: