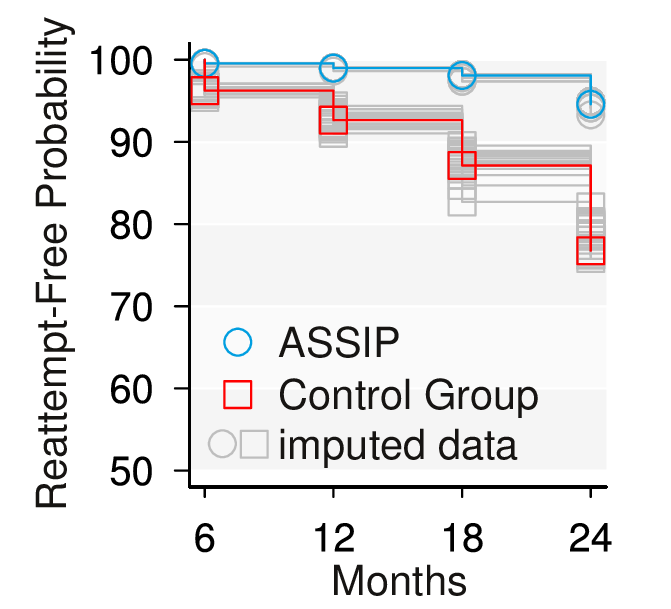
This repository contains data and analyses from the ASSIP 24-months follow-up randomized controlled study.
We provide a short analysis script (survival.R) to replicate the main findings of our study using the original data (assip.RData). Results in our paper may slightly differ because these are based on twenty imputed dataset. Variable description and statistical output see below.
Gysin-Maillart A, Schwab S, Soravia LM, Megert M, & Michel K (2016). A Novel Brief Therapy for Patients Who Attempt Suicide: A 24-months follow-up randomized controlled study of the Attempted Suicide Short Intervention Program (ASSIP). PLoS Medicine 13(9):e1001968. 10.1371/journal.pmed.1001968.
Article in The Washington Post.
amb_sess_t1_6M Number of outpatient sessions at time one (t1), including the last 6 months
bdisum2 BDI (Beck Depression Intventory) sum, after 6 months (t2)
bdisum3 BDI (Beck Depression Intventory) sum, after 12 months (t3)
bdisum4 BDI (Beck Depression Intventory) sum, after 18 months (t4)
bdisum5 BDI (Beck Depression Intventory) sum, after 24 months (t5)
BSS_T2 BSS (Beck Scale for Suicide Ideation) mean, after 6 months (t2)
BSS_T3 BSS (Beck Scale for Suicide Ideation) mean, after 12 months (t3)
BSS_T4 BSS (Beck Scale for Suicide Ideation) mean, after 18 months (t4)
BSS_T5 BSS (Beck Scale for Suicide Ideation) mean, after 24 months (t5)
ITT Intention to treat
repeater_t2 repeated suicide attempts (patient), 1-6 months, dichotomous(0, minimum 1 suicide attempt)
repeater_t3 repeated suicide attempts (patient), 6-12 months, dichotomous(0, minimum 1 suicide attempt)
repeater_t4 repeated suicide attempts (patient), 12-18 months, dichotomous(0, minimum 1 suicide attempt)
repeater_t5 repeated suicide attempts (patient), 18-24 months, dichotomous(0, minimum 1 suicide attempt)
stat_days_t1 Inpatient days at baseline (6 months backward)
sum_stat_12months Sum of inpatient days after 12 months
sum_stat_24months Sum of outpatient days after 24 months
> fit = survfit(Surv(time, repeater) ~ group, type="kaplan-meier")
> summary(fit)
Call: survfit(formula = Surv(time, repeater) ~ group, type = "kaplan-meier")
65 observations deleted due to missingness
group=ASSIP & ASSIP Drop out
time n.risk n.event survival std.err lower 95% CI upper 95% CI
6 229 1 0.996 0.00436 0.987 1
12 170 1 0.990 0.00727 0.976 1
18 111 1 0.981 0.01143 0.959 1
24 56 2 0.946 0.02671 0.895 1
group=CG & CG Drop out
time n.risk n.event survival std.err lower 95% CI upper 95% CI
6 186 7 0.962 0.0140 0.935 0.990
12 134 5 0.926 0.0207 0.887 0.968
18 84 5 0.871 0.0308 0.813 0.934
24 42 5 0.768 0.0513 0.673 0.875
>
> # group difference (Mantel-Haenszel) ----
> survdiff(Surv(time, repeater) ~ group, rho=0)
Call:
survdiff(formula = Surv(time, repeater) ~ group, rho = 0)
n=415, 65 observations deleted due to missingness.
N Observed Expected (O-E)^2/E (O-E)^2/V
group=ASSIP & ASSIP Drop out 229 5 15.2 6.83 16.1
group=CG & CG Drop out 186 22 11.8 8.78 16.1
Chisq= 16.1 on 1 degrees of freedom, p= 5.99e-05
For hazard ratio see exp(-coef) below
> # Cox hazard for discrete data ----
> hazard <- coxph(Surv(time, repeater) ~ group, ties="exact")
> summary(hazard)
Call:
coxph(formula = Surv(time, repeater) ~ group, ties = "exact")
n= 415, number of events= 27
(65 observations deleted due to missingness)
coef exp(coef) se(coef) z Pr(>|z|)
groupCG & CG Drop out 1.7826 5.9451 0.5006 3.561 0.000369 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
exp(coef) exp(-coef) lower .95 upper .95
groupCG & CG Drop out 5.945 0.1682 2.229 15.86
Rsquare= 0.04 (max possible= 0.421 )
Likelihood ratio test= 16.75 on 1 df, p=4.268e-05
Wald test = 12.68 on 1 df, p=0.0003695
Score (logrank) test = 16.11 on 1 df, p=5.992e-05
In the traditional survival analysis an event is generally associated with "death", and only one event is possible per subject (non-recurring events). However, recurring events relax this assumption and are also widely used in the literature, for example multiple relapses from remission for leukemia patients, repeated heart attacks, recurrence of bladder cancer tumors, or deteriorating episodes of visual acuity (Kleinbaum & Klein, 2005). Recurring analysis can be seen as repeated measures analysis, however one issue is that observations are not completely independent. Therefore, we also performed the analysis using non-recurring events.
E='Suizidversuch (mind. 1)'
time=rep(NA,120)
event=rep(NA,120)
for (i in 1:120) {
f = c(mydata$repeater_t2[i]==E, mydata$repeater_t3[i]==E, mydata$repeater_t4[i]==E, mydata$repeater_t5[i]==E)
if (sum(f, na.rm = T) > 0) { # time to event
time[i]=which(f)[1]*6; event[i]=1
} else { # censoring
c=tail(which(!f), n=1)
if (length(c) > 0) {time[i]=c*6; event[i]=0}
else {time[i]=0; event[i]=0}
}
}
> fit = survfit(Surv(time, event) ~ mydata$ITT, type="kaplan-meier")
> summary(fit)
Call: survfit(formula = Surv(time, event) ~ mydata$ITT, type = "kaplan-meier")
mydata$ITT=ASSIP & ASSIP Drop out
time n.risk n.event survival std.err lower 95% CI upper 95% CI
6 59 1 0.983 0.0168 0.951 1.000
12 58 1 0.966 0.0236 0.921 1.000
18 54 1 0.948 0.0291 0.893 1.000
24 53 2 0.912 0.0374 0.842 0.989
mydata$ITT=CG & CG Drop out
time n.risk n.event survival std.err lower 95% CI upper 95% CI
6 53 7 0.868 0.0465 0.781 0.964
12 44 3 0.809 0.0545 0.709 0.923
18 37 4 0.721 0.0637 0.607 0.858
24 30 2 0.673 0.0680 0.552 0.821
> fit = survfit(Surv(time, event) ~ mydata$ITT, type="kaplan-meier")
> summary(fit)
Call:
survdiff(formula = Surv(time, event) ~ mydata$ITT, rho = 0)
N Observed Expected (O-E)^2/E (O-E)^2/V
mydata$ITT=ASSIP & ASSIP Drop out 60 5 12.01 4.09 10.1
mydata$ITT=CG & CG Drop out 60 16 8.99 5.47 10.1
Chisq= 10.1 on 1 degrees of freedom, p= 0.00148
> hazard <- coxph(Surv(time, event) ~ mydata$ITT, ties="exact")
> summary(hazard)
Call:
coxph(formula = Surv(time, event) ~ mydata$ITT, ties = "exact")
n= 120, number of events= 21
coef exp(coef) se(coef) z Pr(>|z|)
mydata$ITTCG & CG Drop out 1.5294 4.6156 0.5228 2.926 0.00344 **
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
exp(coef) exp(-coef) lower .95 upper .95
mydata$ITTCG & CG Drop out 4.616 0.2167 1.657 12.86
Rsquare= 0.082 (max possible= 0.71 )
Likelihood ratio test= 10.22 on 1 df, p=0.001386
Wald test = 8.56 on 1 df, p=0.003436
Score (logrank) test = 10.11 on 1 df, p=0.001478