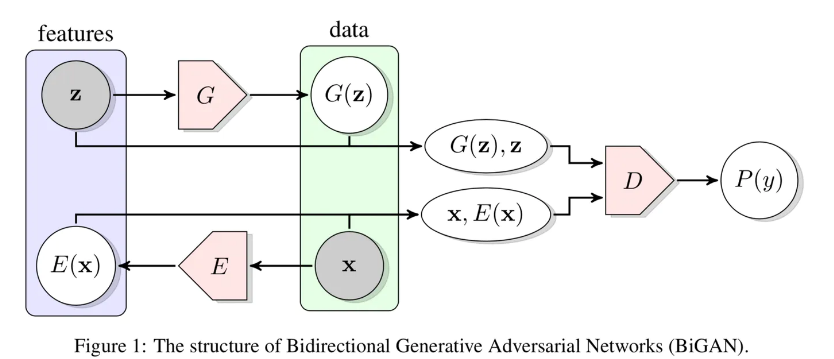
This repository was created as a lean, easy to use & adapt implementation of the 2018 paper
This repository was built in Anaconda (Python 3.7) on Ubuntu 18.04. It is highly recommended you use Anaconda to rebuild this repository as it will ensure
your environment will be solved in terms of dependencies such as CUDA/cuDNN for GPU use.
You have 2 ways to build this environment (both using conda environment):
$ conda create --name <your_venv> --file requirements.txt
or
$ conda env update -n <your_venv> --file environment.yml
- bigan_model.py - includes all the part needed to build and initialize a BiGAN model
- bigan_trainer.py - code for the training the BiGAN, saving checkpoints and visualizations to TensorBoard
- bigan_eval.py - code for running inference on the trained BiGAN with the logic implementated to use it as an anomaly detector + metric visualizations
My specific use case for the EGBAD was for large 4K images, which were needed to be sliced and labeled to be fed to the BiGAN efficiently.
So this module expects large images + json annotations in the same directory like so:
data_dir/
images/img_*.png annotations/img_*.png.json
The module creates a generator which yields batches of (image_batch, label_batch) where the label is whether there's an anomaly or not (relevant only for testing)
It may not be relevant for your use-case, but I've included it anyway so you can better understand the project's flow.
Bottom line is you only need to create a generator yielding (image_batch, label_batch) in order for the train/test to work.
There are some recommendations however:
- input shape should be -
(32, 32, c)or(64, 64, c).
Anything bigger would be very difficult to train. - inputs should be normalized to
[-1 , 1]
You have tools in the repo in the shape of flagscrop_size,resize, and acenter_and_scalefunction to help you prepare the data in such a way.
These are the runner scripts for the trainin phase / inference phase respectivly. You can understand the API and parameter configurations through the abseil app flags listed below the imports. Some of the flags are meant for the data preparation logic of my "slice and label" use case, so feel free to alter it for your use.