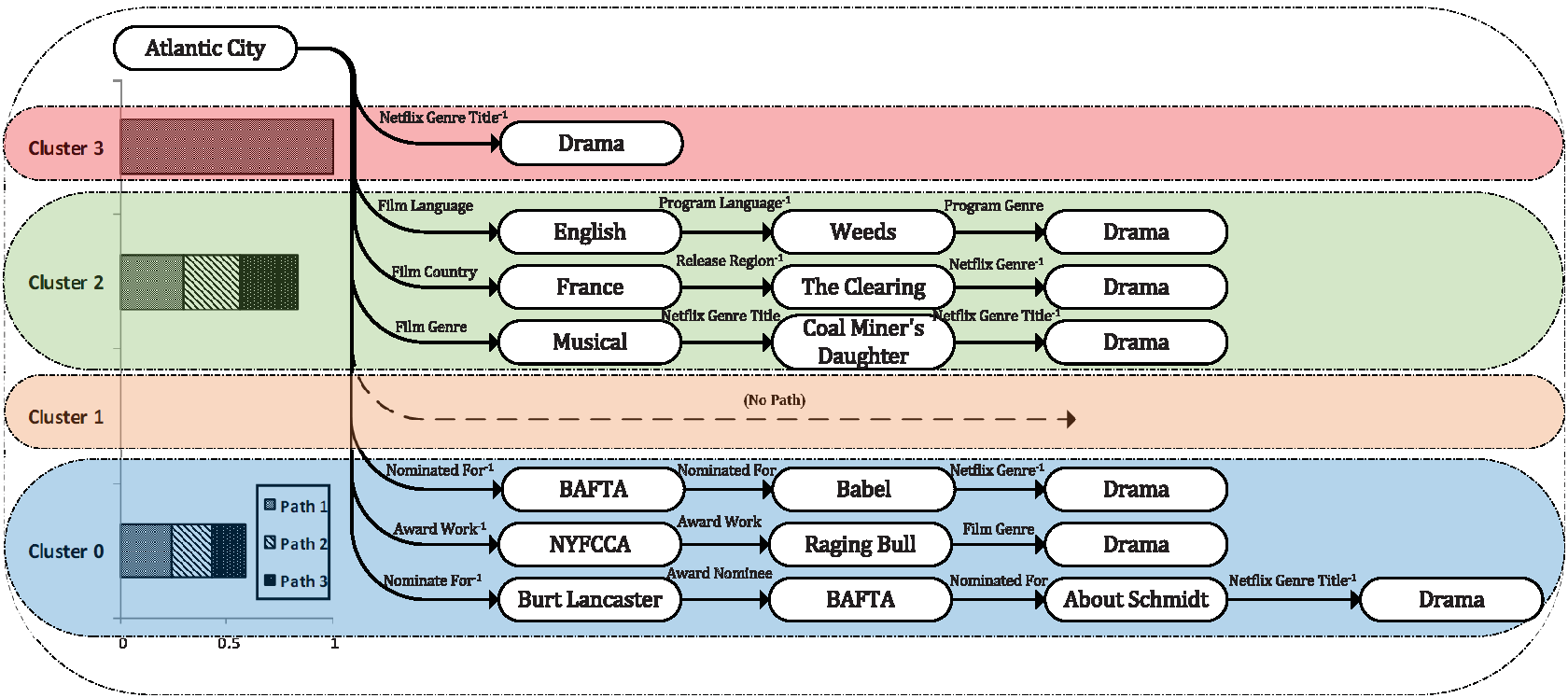
This repository contains code and data for AAAI 2023 paper Multi-Aspect Explainable Inductive Relation Prediction by Sentence Transformer. In the paper, we propose a novel approach for inductive relation prediction namely KRST. KRST adopts sentence transformer for path-based prediction and could provide a comprehensive explanation.
- python 3.6.0
- colorama 0.4.4
- matplotlib 3.3.4
- networkx 2.5.1
- numpy 1.18.5
- scikit_learn 1.1.2
- torch 1.9.1
- tqdm 4.64.0
- transformers 4.18.0
We provide an all-in-one file generate_train_and_test.sh to automatically extract paths, train model, and test model. To specify datasets and few-shot settings, change variables in generate_train_and_test.sh (line 1-3) as follows:
dataset: Available datasets includeFB15k-237-subset,NELL-995-subset,WN18RR-subset,FB15k-237-subset-inductive,NELL-995-subset-inductive, andWN18RR-subset-inductive.suffix: This is for few-shot settings, including_full,_2000, and_1000.finding_mode: whetherheadortailis fixed.
Then by executing the following command:
./generate_train_and_test.shyou can reproduce the confidence results mentioned in the paper. And testing results are stored in save\$dataset$$suffix$\relation_prediction_$finding_mode$\ (see KRST testing section for more details).
Also, you can just download the model trained by our team here and test them using
./test.shAll datasets mentioned in the paper are provided in data\data. They are from BERTRL and are reorganized. If you want to manually create a new dataset, please provide the following files:
train$suffix$.txt: file for training. Each line is a triplet (h,r,t) seperated by '\t'.valid.txtandtest.txt: files for validation and testing.relation2text.txtandentity2text.txt: description file. Each line contains a relation (or entity) and its corresponding descriptions, separated by '\t'.ranking_head.txtandranking_tail.txt(optional): files of positive and negative testing triplets. If not provided, use the following command to generate:
python neg_sampling.py --dataset $dataset --suffix $suffix --finding_mode $finding_mode --training_mode test --neg_num 50After preparing the dataset, you can do path finding and path filtering using the shell file 'generate.sh'. The parameters that can be fine-tuned include:
npaths_ranking: number of paths generated for each triplet.support_type: choice of path filtering function. 0: none, 1: relation path coverage 2: relation path confidence.support_threshold: threshold of path filtering function.search_depth: breadth-first search depth.
Then you can see the generated files in `data\relation_prediction_path_data$dataset$\ranking_$finding_mode$$suffix$'. If you want to skip this step, please download the generated files here.
Using file 'train.sh', you can format sentences, load pre-trained sentence transformer, and train KRST by yourself. The parameters that can be fine-tuned include:
epochs: upper epoch limit.batch_size: batch size.learning_rate: learning rate.neg_sample_num_train,neg_sample_num_validandneg_sample_num_test: number of negative samples (should be equal to generated by negative_sampling.py)max_path_num: number of paths loaded for each triplet (you can load a smaller number of paths than generated by path finding).
After training, the best model is saved in save\$dataset$$suffix$\relation_prediction_$finding_mode$\best_val.pth.
By default, KRST loads the saved best validation model for testing. Run 'test.sh' to get the test results (testing results are stored in save\$dataset$$suffix$\relation_prediction_$finding_mode$\):
scores.txt: each line contains the scores for one positive triplet and corresponding negative triplets. Scores are separated by '\t' and follow the order inranking_$finding_mode$.indexes.txt: each line contains the sorted index of scores (argsort in python) inscores.txt.
You can just download the model trained by our team here and test them.
Before doing interpretation, you need to put two files in data\relation_prediction_path_data\$dataset$\interpret:
best_val.pth: model loaded for interpretation.interpret.txt: triplets to be interpreted.
Then run 'interpret.sh' to generate clustering results (pyfig.eps) and confidence score (result.txt). You can download an example from here.
Please refer to citations here.
For any questions, feel free to leave an issue. Thank you very much for your attention and further contribution :)