Comments (3)
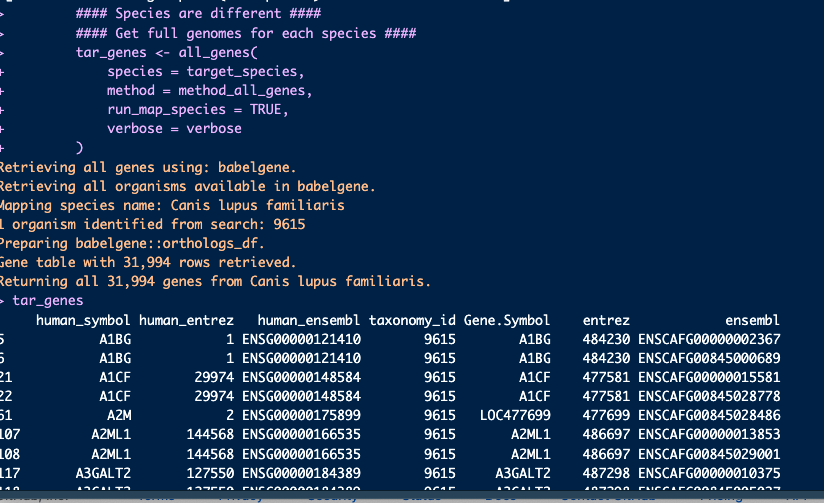
Noticing that this is returning data where the Gene.Symbol is actually the human ortholog, not the dog gene name as we would expect:
tar_genes <- all_genes(
species = "Canis lupus familiaris",
method = "babelgene",
run_map_species = TRUE)from orthogene.
I think this might be the root of it:
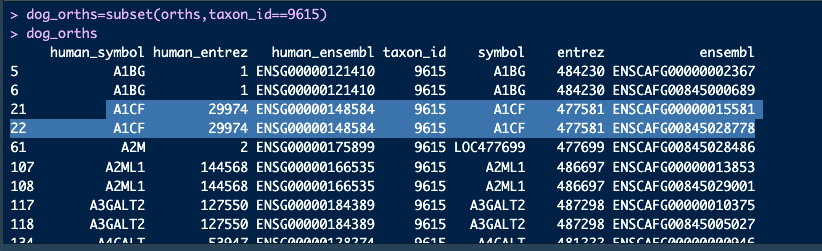
dog_orths=subset(babelgene:::orthologs_df,taxon_id==9615)
dog_orthsWe can see that many genes (e.g. A1BG, A1CF) have the same gene symbol across species and are 1:1 in that sense. But there are multiple mappings from the dog symbol to the dog ensembl ID. This makes orthogene think theyre not 1:1 orthologs and thus drops them. But if all we care about is gene symbol to gene symbol, then we should keep them.
from orthogene.
This was exactly the issue! Fixed it by adding an extra step to drop_non121
if(isTRUE(symbol_only)){
gene_map <- data.table::as.data.table(gene_map)
if("support_n" %in% names(gene_map)){
data.table::setorderv(gene_map,"support_n",-1)
}
gene_map <- gene_map[,.SD[1],by=c("input_gene","ortholog_gene")]
}
After rerunning report_orthologs, you get a near-perfect reconstruction of species similarity to humans. The exception being that dogs now appear more closely related to humans than mice (which is technically wrong, but close enough given that we're only looking at protein coding genes with 1:1 orths):

from orthogene.
Related Issues (20)
- Look into OMA
- Add phylogenetic tree databases
- Scale silhouettes HOT 2
- How to find all supported species? HOT 1
- How to use older annotations with convert_orthologs method? HOT 2
- Issue with create_background - found in EWCE HOT 4
- babelgene tests HOT 1
- Add earthworm orthologs HOT 1
- `Matrix.utils` deprecated HOT 1
- Dependency "Matrix.utils" is no longer available HOT 2
- `rphylopic` failing HOT 3
- Replace `ggtree`/`ggimage` HOT 1
- GHA: `Package suggested but not available: ‘rworkflows’` HOT 1
- GHA: Docker: Error: both username and password must be set to login both username and password must be set to login HOT 2
- Consider adding ortholog mappings from Zoonomia
- convert_orthologs() output is list, with "Warning: Coercing LHS to a list" HOT 5
- Consider integrating `taxize`
- Determine which journal to publish in
- The result was 0 on R package orthogene HOT 6
Recommend Projects
-
 React
React
A declarative, efficient, and flexible JavaScript library for building user interfaces.
-
Vue.js
🖖 Vue.js is a progressive, incrementally-adoptable JavaScript framework for building UI on the web.
-
 Typescript
Typescript
TypeScript is a superset of JavaScript that compiles to clean JavaScript output.
-
TensorFlow
An Open Source Machine Learning Framework for Everyone
-
Django
The Web framework for perfectionists with deadlines.
-
Laravel
A PHP framework for web artisans
-
D3
Bring data to life with SVG, Canvas and HTML. 📊📈🎉
-
Recommend Topics
-
javascript
JavaScript (JS) is a lightweight interpreted programming language with first-class functions.
-
web
Some thing interesting about web. New door for the world.
-
server
A server is a program made to process requests and deliver data to clients.
-
Machine learning
Machine learning is a way of modeling and interpreting data that allows a piece of software to respond intelligently.
-
Visualization
Some thing interesting about visualization, use data art
-
Game
Some thing interesting about game, make everyone happy.
Recommend Org
-
Facebook
We are working to build community through open source technology. NB: members must have two-factor auth.
-
Microsoft
Open source projects and samples from Microsoft.
-
Google
Google ❤️ Open Source for everyone.
-
Alibaba
Alibaba Open Source for everyone
-
D3
Data-Driven Documents codes.
-
Tencent
China tencent open source team.

from orthogene.